You probably need to provide more details on what you are exactly doing.
With the following script
import harp
import numpy as np
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
def harp_l3meshplot(product, value, colorrange=None, colortable='jet'):
variable = product[value]
gridlat = np.append(product.latitude_bounds.data[:,0], product.latitude_bounds.data[-1,1])
gridlon = np.append(product.longitude_bounds.data[:,0], product.longitude_bounds.data[-1,1])
if colorrange is not None:
vmin, vmax = colorrange
else:
vmin = np.nanmin(variable.data)
vmax = np.nanmax(variable.data)
fig=plt.figure(figsize=(20, 10))
ax = plt.axes(projection=ccrs.PlateCarree())
img = plt.pcolormesh(gridlon, gridlat, variable.data[0,:,:], vmin=vmin, vmax=vmax,
cmap=colortable, transform=ccrs.PlateCarree())
ax.coastlines()
ax.gridlines()
cbar = fig.colorbar(img, ax=ax, orientation='horizontal', fraction=0.04, pad=0.1)
cbar.set_label(f'{variable.description} [{variable.unit}]')
cbar.ax.tick_params(labelsize=14)
plt.show()
files = [
"/Users/sander/Downloads/S5P_OFFL_L2__CH4____20210608T040316_20210608T054446_18923_01_010400_20210609T210658.nc",
"/Users/sander/Downloads/S5P_OFFL_L2__CH4____20210608T054446_20210608T072616_18924_01_010400_20210609T223439.nc",
]
operations = ";".join([
"CH4_column_volume_mixing_ratio_dry_air_validity>50",
"latitude > -45",
"latitude < -10.2",
"longitude > 110.6",
"longitude < 157",
"bin_spatial(1739,-45,0.02,2320,110.6,0.02)",
"keep(CH4_column_volume_mixing_ratio_dry_air,cloud_fraction,latitude_bounds,longitude_bounds)",
])
post_operations="bin();squash(time, (latitude_bounds, longitude_bounds))"
grid = harp.import_product(files, operations, post_operations=post_operations)
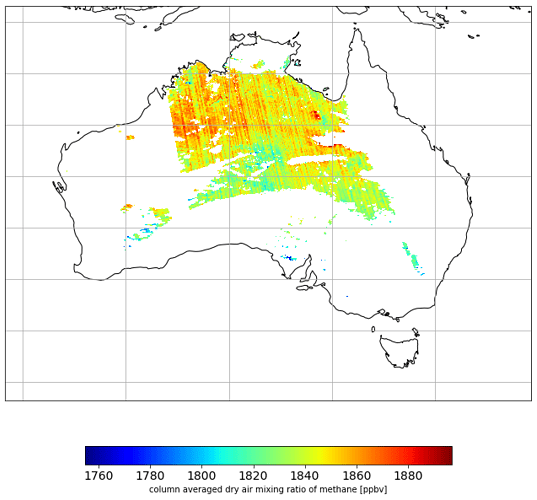
harp_l3meshplot(grid, "CH4_column_volume_mixing_ratio_dry_air")
I am getting the following plot:
Which looks fine to me